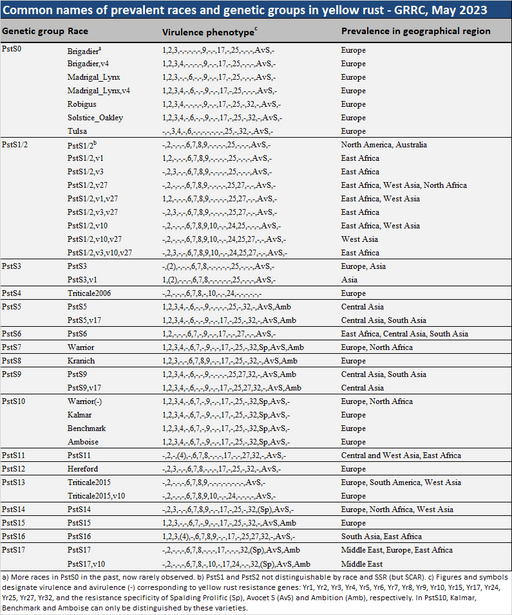
Definitions of races and genetic groups
Definition of race
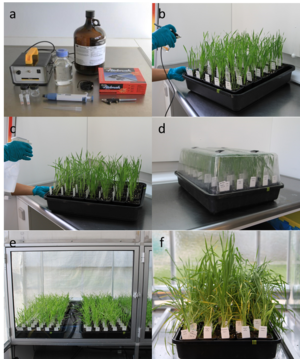
Race is defined by the pattern of compatible and incompatible interactions between host and pathogen. The pathogen phenotype is ‘virulent’ in case of a compatible interaction, shown by ‘high’ infection type scores on host differential lines, and it is ‘avirulent’ in case of incompatible interactions conferred by ‘low’ infection types (Fig. 1). The genetic interpretation of such race (pathotype) data depend on the extent that R-genes have been identified in the host differential lines, and additional resistance specificities have been resolved by exposure of the differential lines to a wide array of pathogen isolates of diverse geographical and evolutionary origin.
The virulence phenotype is shown by figures and symbols designating virulence and avirulence (-) corresponding to yellow rust resistance genes: Yr1, Yr2, Yr3, Yr4, Yr5, Yr6, Yr7, Yr8, Yr9, Yr10, Yr15, Yr17, Yr25, Yr27, Yr32, and the resistance specificity of Spalding Prolific (Sp), and Ambition (Amb), respectively. All prevalent races except Triticale2006 (PstS4) were virulent on Avocet S.
The yellow rust race phenotyping methodology used in the GRRC lab is described in detail in Hovmøller et al. 2017: Wheat Rust Diseases: Methods and Protocols, ed. S. Periyannan. Springer, p. 29-40. Download accepted for non-commercial and educational purposes.
Definition of common names for races and genetic groups
Races associated with major rust epidemics have been assigned a common name according to their genetic group defined by molecular genotype (Ali et al., 2017). Each group consists of one or more closely related races, named Pst followed by a forthcoming digit. Race variants within genetic groups are named according to the name of the group followed by the additional virulence(s), separated by comma. Genetic groups and race names are indicated with normal/bold and Italic/normal characters respectively.
Races in the NW-European population, which have previously been assigned a common race name according to the host varieties where they first caused epidemics, were part of a single clonal group termed PstS0. The PstS1 and PstS2 lineages represented two closely related groups of aggressive isolates defined by AFLP, microsatellite and SCAR markers.
The groups PstS3 and onwards are defined by microsatellite genotype. PstS3 represents a clonal group prevalent in southern Europe, North Africa and West Asia, and PstS4 consists of races prevalent on triticale in Northern Europe. PstS5, PstS6 and PstS9 consisted of races prevalent in Central Asia and East Africa. Races in the PstS7 and PstS8 genetic groups are known by the farming community in Europe as Warrior and Kranich, whereas PstS10 represents a group currently containing one race, Warrior(-), first detected in Europe in 2012 and prevalent in most parts of Europe since 2014.
The documentation for PstS11 and onwards is not yet published. PstS11 was first detected in Afghanistan in 2012, thereafter spreading to additional countries in Central Asia and East Africa. PstS12 consists of the Hereford race detected in Sweden, PstS13 consists of the Triticale2015 race, which is prevalent on triticale and durum wheat on several continents. PstS14 consists of a single race, first detected in 2016 in North Africa and Europe. PstS15, consisting of a single race, has only been detected in Europe. PstS16 was first detected in South Asia, but since 2020 also detected in East Africa.
References
- Hovmøller et al. 2017: Race Typing of Puccinia striiformis on Wheat. In: Wheat Rust Diseases: Methods and Protocols, ed. S. Periyannan. Springer, p. 29-40. Downloading accepted for non-commercial and educational purposes.
- Ali et al. 2017: Yellow Rust Epidemics Worldwide Were Caused by Pathogen Races from Divergent Genetic Lineages, Frontiers in Plant Science 8, June2017. https://doi.org/10.3389/fpls.2017.01057.