Potato blight mapped in Europe - 2014
A team of researchers tracking the 2014 population of the potato late blight pathogen have added to the 2013 data to extend the spatial diversity plots and combine this with novel genetic analysis tools that visualise the distribution and diversity of dominant clones and reveals novel genetically diverse isolates in some regions.
Since its arrival in the nineteenth century, Phytophthora infestans, the cause of late blight has remained a serious threat to potato crops. Although we are now better equipped to control the disease than in the past, an evolving pathogen population continues to challenge our management practices. Rapid changes in P. infestans populations causing late blight in Europe, America and Asia, including the emergence of strains with altered pathogenicity or reduced fungicide sensitivity have been observed. Monitoring of populations and characterization of invasive genotypes help optimise IPM strategies, as required by EU Directive 2009/128/EC on the sustainable use of plant protection products. The changes in P. infestans populations directly influence the development and deployment of resistant cultivars, the performance of disease warning systems and the efficacy of plant protection products. Coordinated and continuous pathogen monitoring was proposed by the EuroBlight consortium at its meeting in 2013. Here, researchers in the Netherlands, Scotland and Denmark, working with partners from research labs and industry, present their second report on its pathogen monitoring in potato crops in 2014.
The project hinged on the distribution of ‘FTA cards’ to hundreds of disease ‘scouts’ from across the industry who visited blight-infected crops. Disease lesions were pressed on the cards and returned to the laboratories where the pathogen DNA was fingerprinted at Wageningen University and Research Centre and the James Hutton Institute. Disease pressure was generally high in 2014 and 1552 samples were genotyped. This data also includes that from the AHDB Potato Council ‘Fight Against Blight’ campaign in British crops. The fingerprint patterns of all the P. infestans samples were combined and compared to those found previously. In 2014 the team have extended from mapping to include population genetic analysis that uses an R-based tool POPPR linked to the pathogen diversity database by the team at Arhus University.
| A structure within the population is apparent with a broad distribution of dominant clones to the west of Europe and a scatter of novel, genetically diverse isolates defined in a category termed ‘other’ to the east (Figure 1). The aggressive clone EU_13_A2 (blue-13) comprised 36% of the population and was present from the Canaries to Norway and as far east as Romania. It is resistant to phenylamide fungicides and has been dominant in European populations for some years. Growers should be aware that this clone is more difficult to manage than others. Another aggressive clone termed EU_6_A1 (pink-6) made up 32% of overall population, was dominant in Great Britain, present in France and Belgium and recorded once in Poland. Two older clones were found at lower frequencies; EU_1_A1 (3.6%) clustering in Belgium and northern France and EU_8_A1 (3.7%) only in the UK. |
|
| Figure 1. Genotype map. Go to live map. |
The genetically diverse ‘Other’ samples made up 23% of the samples and although they were found in many regions they were recovered at the highest frequency in the east and north-east of Europe (See Genotype frequency map). In contrast to the clonal types, we do not know what traits these isolates have but each will have originated from soil-borne oospores and the risks from this inoculum may be reduced by using longer rotations.
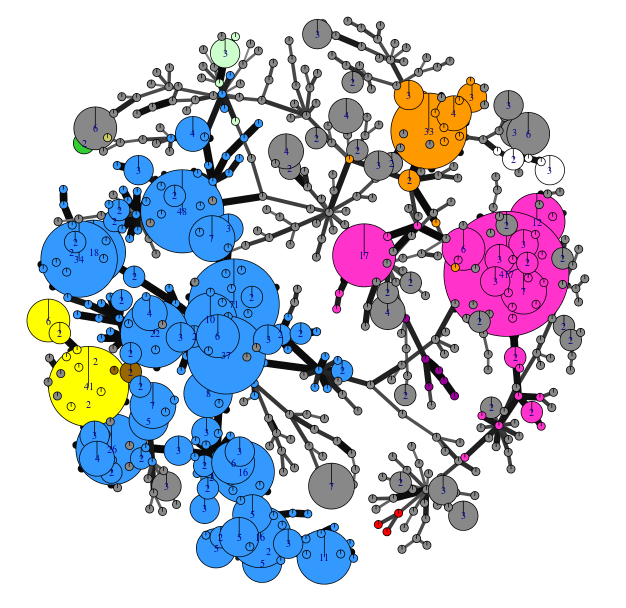
An overview of the genetic diversity data has been generated by a novel link between an analysis tool (POPPR) and the pathogen database (top left figure). Nodes are coloured by clone and show linked sub-clonal clusters of related isolates compared to the more distinct grey ‘other’ genetically diverse isolates. More detailed analysis is underway to examine the year to year trends using these methods. This model of pathogen tracking offers a rapid, cost-effective and co-ordinated approach to understanding pathogen change on a European scale. Data on the dominant clones has been passed to growers, advisors, breeders and agrochemical companies to provide practical management advice and shape longer-term strategies. Furthermore, early warning of newly evolved clones will enable a timely response from the industry.
The Euroblight network is in discussion with other networks in the Americas and Asia and encourages continued co-operation between groups involved in managing late blight to exploit the database and tools for improved awareness and blight management on a global scale.
We will continue the project in 2015 so please make contact with the project team if you would like more information. We thank all the partners who have contributed to the funding and data collection.
Companies and institutions that participated in the sampling and sponsored the project
ADAMA, Agrifirm, Agriphar, BASF SE, Bayer CropScience AG, Bayerische Landesanstalt für Landwirtschaft, Belchim Crop Protection, Centre Wallon de Recherches Agronomiques, Certis, Cheminova, CZAV, Dupont de Nemours, Emsland Group, Germicopa SAS, HZPC Holland B.V., AHDB Potato Council, Neiker, Nordisk Alkali, PCA, Profytodsd, Swedish University of Agricultural Sciences and Syngenta Agro GmbH.
Contacts
Jens G. Hansen, Poul Lassen and Sanmohan Baby, Aarhus University: Potato late blight toolbox, EuroBlight website, Databases and web tools. Contact: jensg.hansen@agro.au.dk
Geert Kessel, Wageningen University and Research Centre. Coordinator of sampling, distribution of FTA cards, SSR analysis and quality control of data. Contact: geert.kessel@wur.nl
David Cooke: James Hutton Institute, Dundee. SSR analysis, collation and control of historical data to be added to the database. Contact: David.Cooke@hutton.ac.uk
Download the PDF version of this news story here
 |  |  |
