Results of the EuroBlight potato late blight monitoring in 2018
EuroBlight is continuously examining the ongoing evolution of the European population of the potato late blight pathogen and now reports on the 2018 results. Approximately 1000 samples were genotyped from 22 countries.
Key findings:
Three emerging clones (37_A2, 36_A2 and 41_A2) increased their combined frequency from 10% in 2016 to 36% of the population in 2018
These more recent clones are displacing the established 13_A2, 6_A1 and 1_A1 clones which reduced from 60 to 40% of the samples over the same period
One fifth of the population comprised ephemeral, genetically diverse isolates consistent with oospore-borne inoculum.
A regional pattern in the dominance of clones versus sexual recombinants was observed across Europe. Some implications of these displacements and ongoing changes are discussed.
The weather in year 2018, was very dry and unfavourable for late blight development. Therefore, the number of samples were lower than normal. Nonetheless, over 1000 samples were genotyped from 22 countries
How did we do it?
Since its arrival in the nineteenth century, Phytophthora infestans, the cause of potato late blight, has remained a serious threat to European potato production. Although we are now better equipped to control the disease than in the past, evolving pathogen populations continue to challenge integrated management practices. Rapid changes in P. infestans populations causing late blight in Europe, America and Asia, including the emergence of strains with increased aggressiveness or reduced fungicide sensitivity, have been observed. Indeed, the changes in P. infestans populations directly influence the development and deployment of resistant cultivars, the performance of disease warning systems and the efficacy of plant protection products.
Therefore, coordinated and continuous pathogen monitoring was proposed by the EuroBlight consortium at its meeting in 2013 and is now implemented as an EU-wide monitoring activity, including all stakeholders. We continue to monitor populations and characterise the invasive genotypes to help optimise IPM strategies, as required by EU Directive 2009/128/EC on the sustainable use of plant protection products.
As in previous years, FTA sampling cards were distributed to ‘scouts’ across the industry, who visited blighted crops and pressed lesions onto the cards. The cards and associated metadata were returned to laboratories at the James Hutton Institute, Dundee and INRA, Rennes for pathogen DNA fingerprinting using simple sequence repeat markers. Comparisons with fingerprint data from previous seasons enabled samples to be allocated to clonal lineages of the pathogen or as new genotypes and the data were uploaded to the EuroBlight database.
Disease pressure in 2018 was relatively low due to the hot dry weather across much of the season. Nonetheless, 17 partner organisations collected 1074 samples from 24 European countries. Included also is data from partners in the IPMBlight2.0 project, which generates pathogen phenotype data to support IPM strategies. The genotype data from 2013-2018 now comprises over 8000 samples from 34 countries. Support of international groups also generated data for parts of Asia, South America and North Africa.
What did we discover?
Over the last six years, 60-79% of the sampled population comprised known clonal lineages that recur each season. The remaining samples were novel, genetically diverse genotypes found at a single location in one season and grouped in a category termed ‘Other’ (Figure 1).
 |  |
|---|---|
| Link to live map |
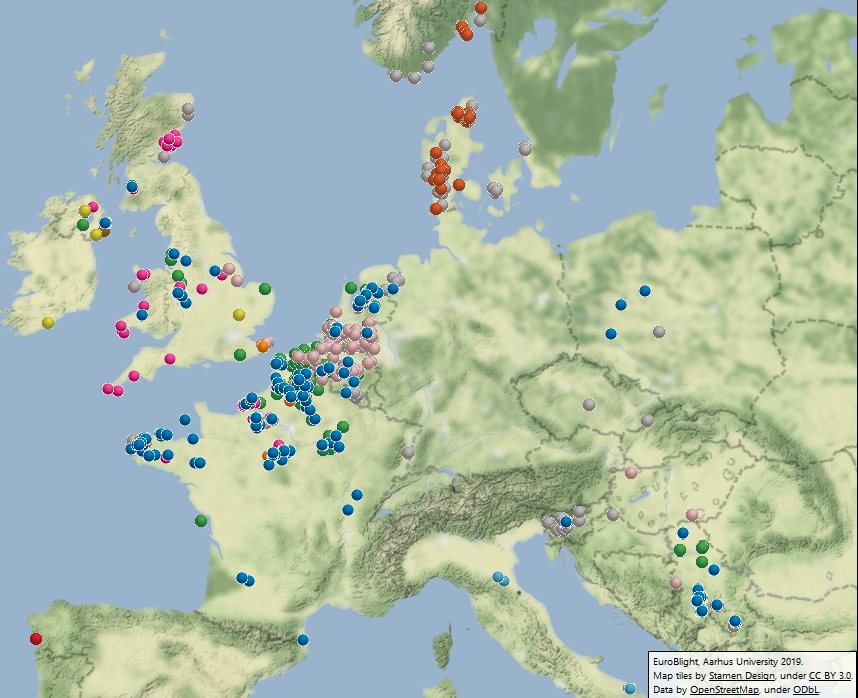
In 2018, clone EU_13_A2 (blue-13) remained the most frequently sampled and widespread genotype with 25% of the samples from 10 countries. The distribution of this aggressive clone EU_13_A2 with its resistance to metalaxyl continues to affect management efficacy in Europe, parts of Asia and North Africa, reinforcing the need for pathogen data to support IPM best practices. The frequency of EU_6_A1 at 13% was the lowest it has been in six years and localised to France and the UK. Similarly, the frequency of EU_1_A1 further decreased from 2.2 to 1.6% of the population. A progressive displacement of these three lineages is occurrin (fig 1)
Despite the warm dry weather, three newer clones (EU_36_A2, EU_37_A2 and EU_41_A2) increased in frequency in 2018. EU_36_A2 was first sampled at low frequencies in the starch potato areas in Germany and the Netherlands in 2014 and had spread across the Netherlands into Belgium, the UK, Denmark and Poland by 2017. In 2018 it was also sampled on crops in Spain, Hungary and Serbia and made up 16% of the samples (up from 10% in 2017). Genotype EU_37_A2 was first detected in Noordoostpolder in the Netherlands in 2013 and had become established in England, Belgium and northern France comprising 14% of the samples by 2017. In 2018 its frequency increased to 16% and its range expanded with first findings in Serbia, Slovenia, Northern Ireland and Scotland. Clone EU_41_A2 was first recorded in Denmark in 2013 and had spread to southern Norway and Sweden and northern Poland by 2017. In 2018 it was recorded in Norway and two regions of Denmark, comprising 18% of the Danish and 4.4% of the European population. The survival and spread of these clones, when others are decreasing or have failed to establish, suggests they are evolutionarily fit and could be more challenging to manage. The insensitivity to fluazinam in EU_37_A2 reported by Wageningen University in 2017 has been confirmed in other studies. Fungicide sensitivity testing in laboratories in Wageningen University and The James Hutton Institute has also shown that EU_36_A2 and EU_37_A2 isolates formed consistently larger lesions than those of the older lineages on leaves at very low dose rates of four key fungicide active ingredients. This supports aggressiveness testing as part of the IPMBlight2.0 project at INRA that revealed that the 36_A2 isolates tested formed large average lesion sizes with abundant sporulation. Such properties are likely to impact on management practices and may explain the spread of these lineages.
Lastly, the genetically diverse ‘Other’ samples comprised 21% of the sampled population in 2018, down slightly from 25% in 2017. However, reduced sampling in the Baltic states and Nordic regions may explain this reduction. The IPMBlight2.0 project is also testing the traits of these ‘Other’ genotype isolates and evaluating European forecast models which is generating data that will help improve late blight management and forecasting in the future.
 |  |
|---|
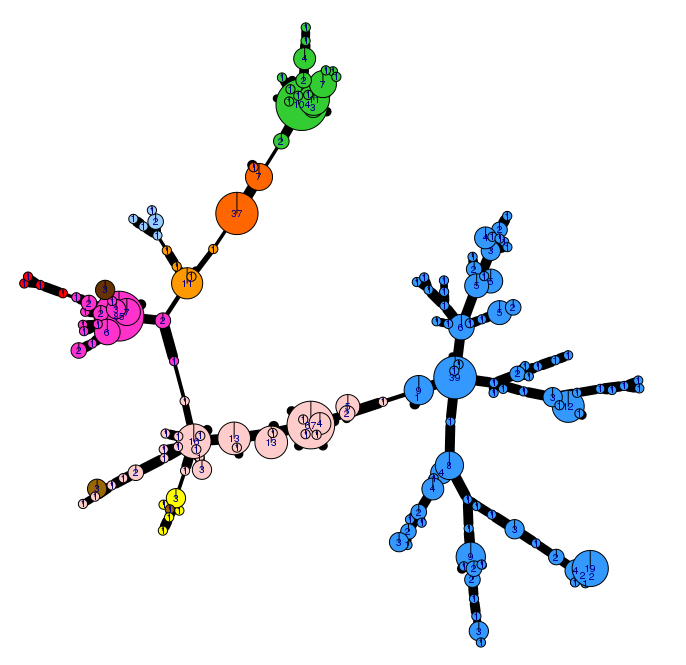
The genetic diversity of the 2018 population has been visualised (Fig. 2) using an analysis tool (poppr 2.0) linked to the EuroBlight pathogen database. The minimum spanning network shows sub-clonal diversity within the 15 year-old EU_13_A2 lineage. The clonal and within-clone variation is being used to track the evolution and spread of these pathogen populations across Europe and beyond. ‘Other’ isolates (not shown) are genetically diverse and distributed across the whole network. Detailed analysis is underway to examine population change using these tools.
The EuroBlight model of pathogen tracking is a rapid, cost-effective and co-ordinated approach to understanding pathogen evolution on a European scale. Data on the dominant clones have been passed to growers, advisors, breeders and agrochemical companies to provide practical management advice and shape longer-term strategies. The data provide an early warning of the incidence and spread of novel clones that is enabling a timely response from the industry.
The EuroBlight network continues to harmonise methods with other networks in the Americas, Asia and Africa and encourages continued co-operation between groups involved in managing late blight to exploit the database and tools for improved awareness and blight management on a global scale. We will continue the project in 2019, so please contact the project team if you would like more information and FTA cards. We thank all the partners who have contributed samples and supported the project.
Companies and institutions that participated in the sampling and sponsored the project (2013-2018)
Aarhus University, ACVNPT, ADAMA, AFBI, Agrifirm, Agricultural Institute of Slovenia, Agriphar, AHDB Potatoes, ARVALIS-Institut du Végétal, BASF SE, Bayer CropScience AG, Bayerische Landesanstalt für Landwirtschaft, Belchim Crop Protection, Centre Wallon de Recherches Agronomiques, Certis, Cheminova, CropSolutions, CUConsulting, CZAV, Delphi, Dupont de Nemours, Emsland Group, Estonian University of Life Sciences, Germicopa SAS, HZPC Holland B.V., INRA, Institute of Plant Protection and Environment in Serbia, The James Hutton Institute, Neiker, Nordisk Alkali, NIBIO, PCA, The Plant Breeding and Acclimatization Institute (IHAR), Profytodsd, Swedish University of Agricultural Sciences, Syngenta Agro GmbH, SynTech Research, TEAGASC and Wageningen University.
Contacts
- Jens G. Hansen & Poul Lassen, Aarhus University: Contact: jensg.hansen@agro.au.dk
- David Cooke and Alison Lees: James Hutton Institute, Dundee. Contact: david.cooke@hutton.ac.uk
- Geert Kessel and Huub Schepers, Wageningen University and Research Centre. Contact: geert.kessel@wur.nl
- Didier Andrivon, Roselyne Corbiere: Contact: Didier.Andrivon@inra.fr
Download the news story as pdf